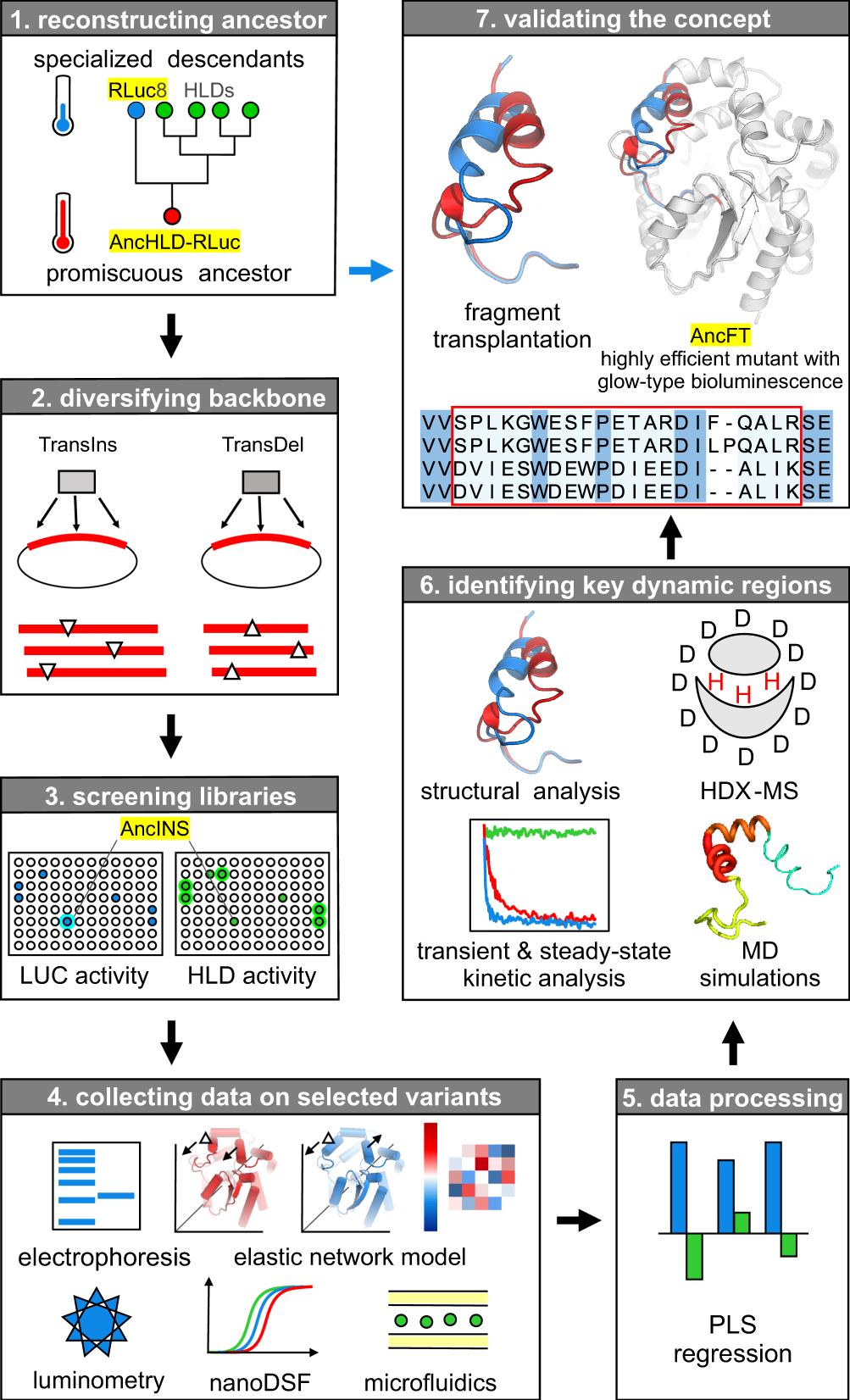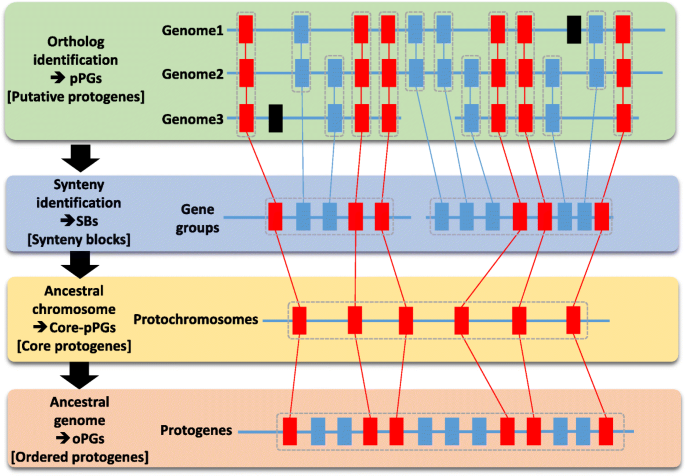
Paleogenomics: reconstruction of plant evolutionary trajectories from modern and ancient DNA | Genome Biology | Full Text

River capture or ancestral polymorphism: an empirical genetic test in a freshwater fish using approximate Bayesian computation | Laboratório de Ictiologia – Universidade Federal do Rio Grande do Sul
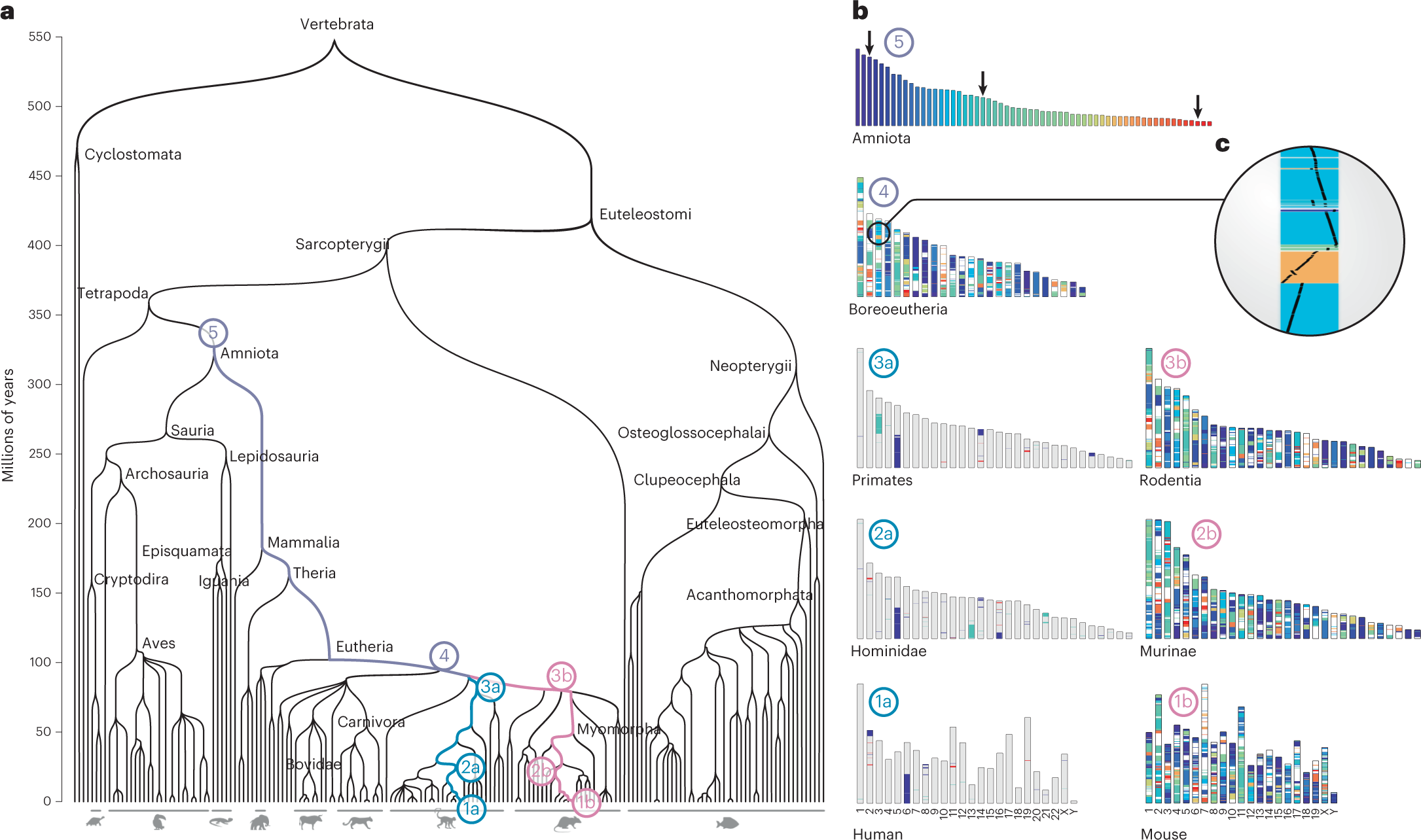
Reconstruction of hundreds of reference ancestral genomes across the eukaryotic kingdom | Nature Ecology & Evolution
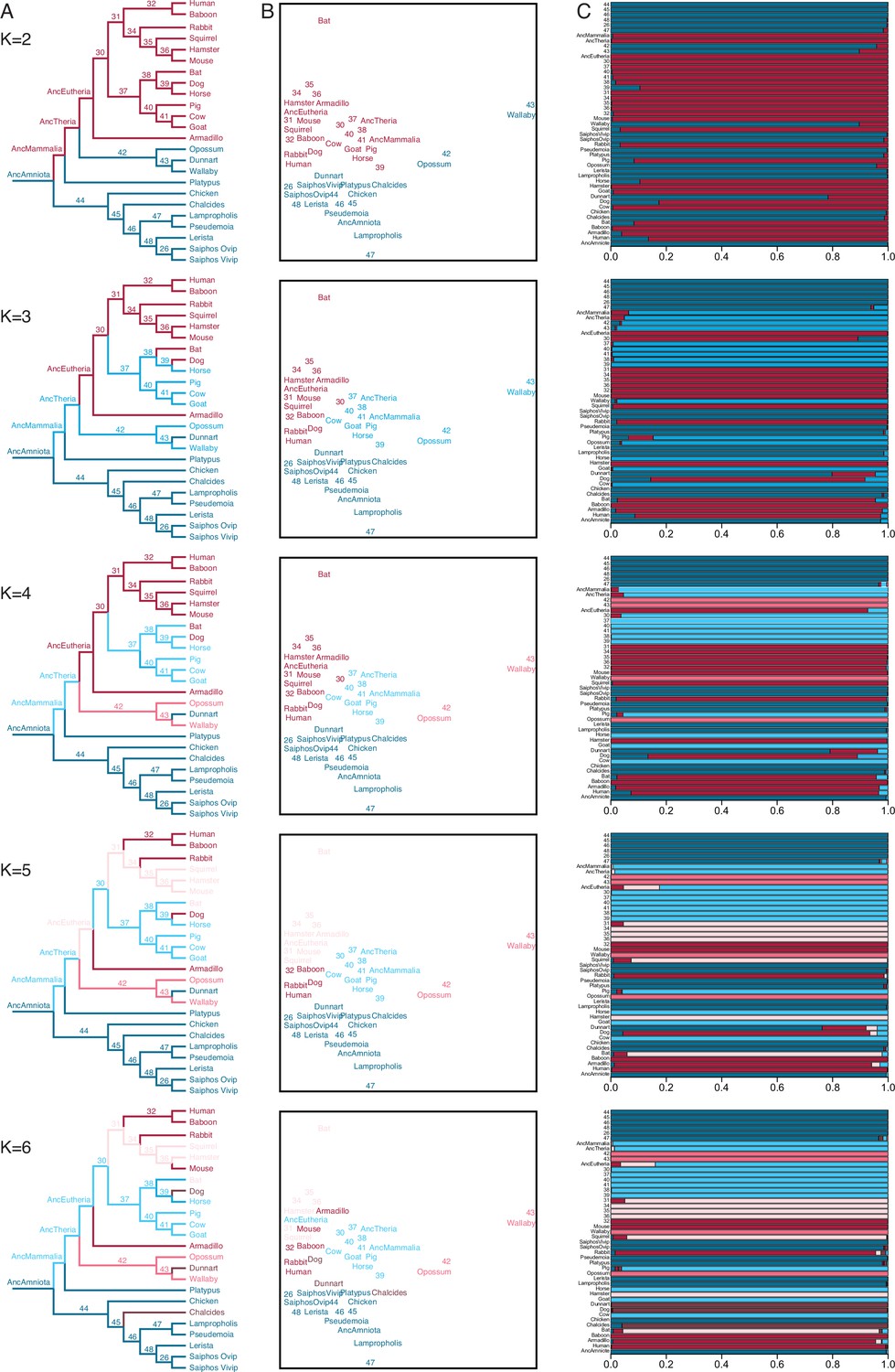
Gene expression phylogenies and ancestral transcriptome reconstruction resolves major transitions in the origins of pregnancy | eLife
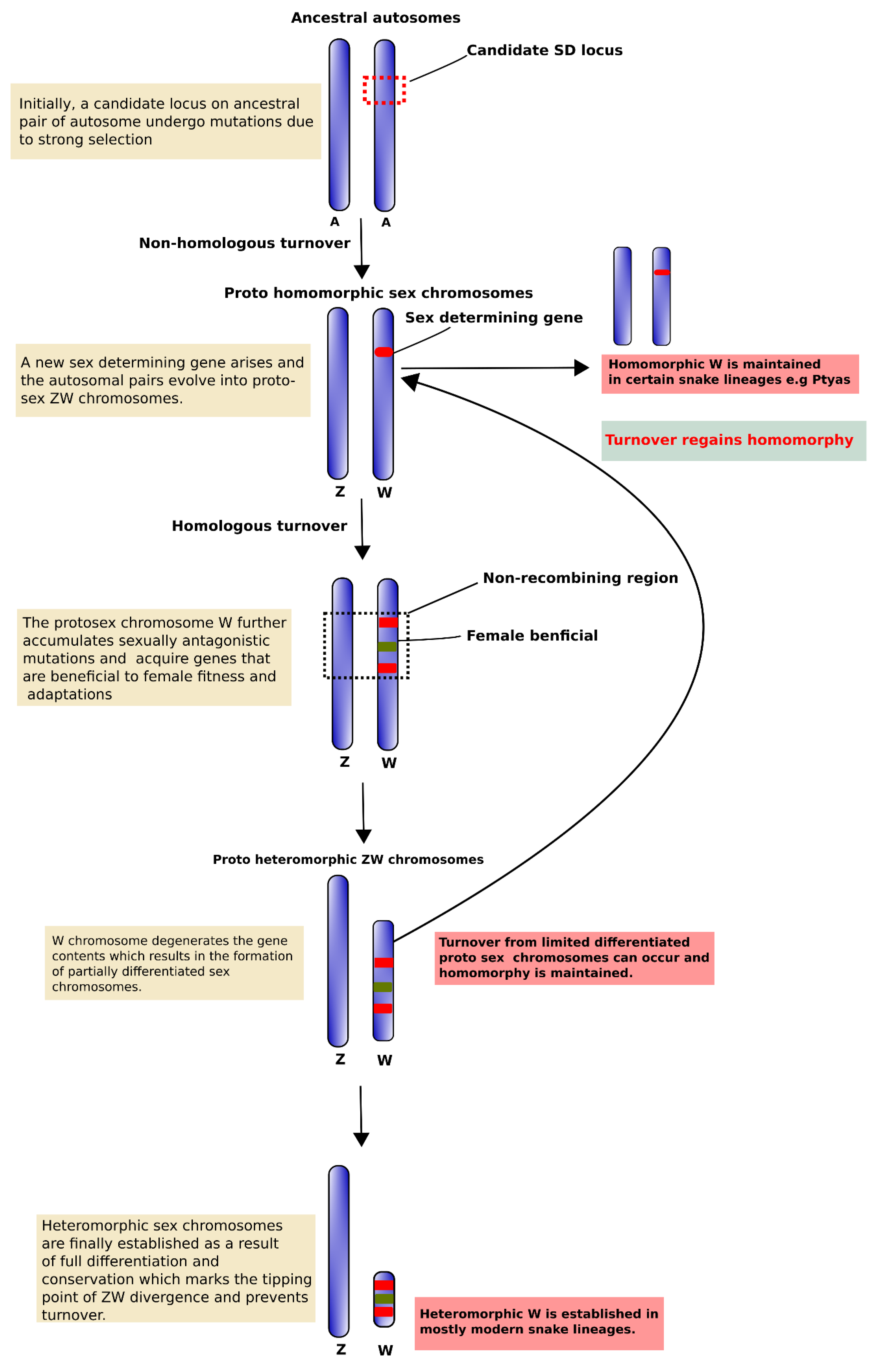
Cells | Free Full-Text | Snake W Sex Chromosome: The Shadow of Ancestral Amniote Super-Sex Chromosome
Mapping gene flow between ancient hominins through demography-aware inference of the ancestral recombination graph | PLOS Genetics

Possible fates of duplicated genes. The most common outcome of gene... | Download Scientific Diagram

Discovery of synthetic lethal and tumor suppressor paralog pairs in the human genome - ScienceDirect

Gerry Tonkin-Hill on Twitter: "Panstripe is available as an R package and requires a core genome phylogeny and gene presence/absence matrix as input. These are output by most pangenome inference tools including
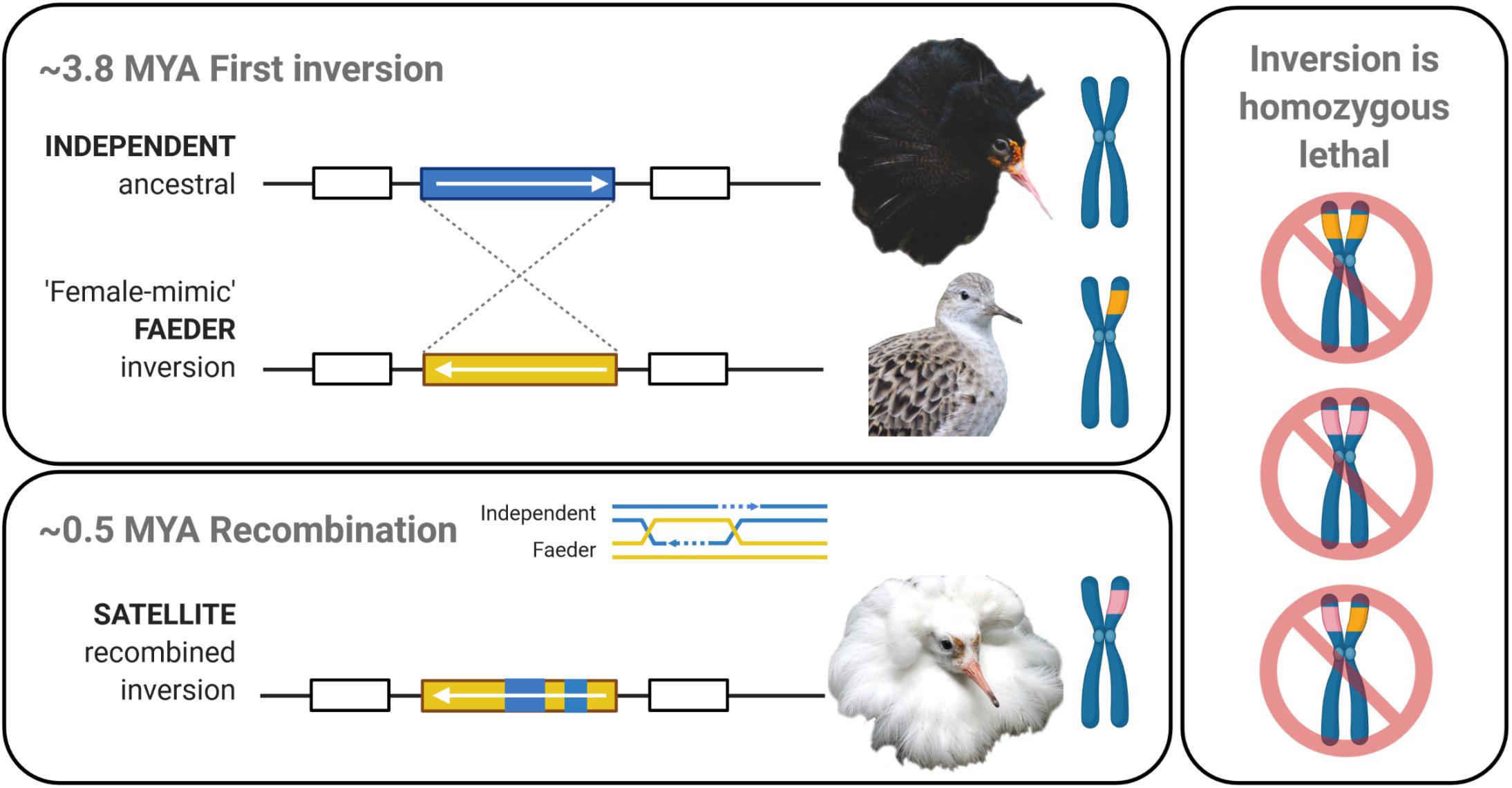
Frontiers | Gene Expression Modification by an Autosomal Inversion Associated With Three Male Mating Morphs


![MK test result for larger pseudoobscura dataset Fuller et. al [31] | Download Table MK test result for larger pseudoobscura dataset Fuller et. al [31] | Download Table](https://www.researchgate.net/publication/327969453/figure/tbl1/AS:676358824919041@1538267918471/MK-test-result-for-larger-pseudoobscura-dataset-Fuller-et-al-31.png)